sunspot <- datasets::sunspot.month
plot(sunspot)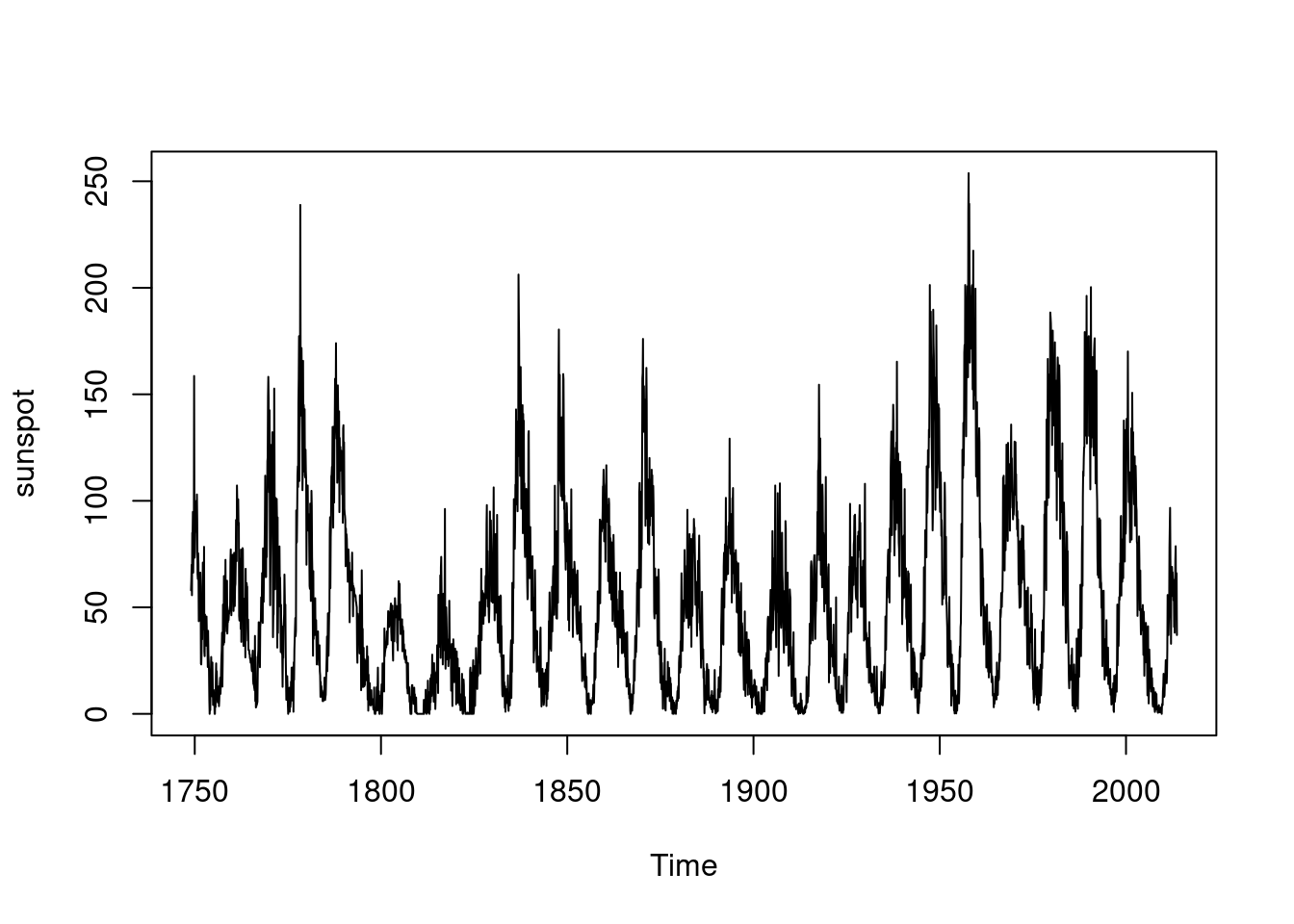
If a time series is “regular”, that is it has equal time steps between observations, and no missing observations, then its power spectrum can be estimated from its Fourier transform.
sunspot <- datasets::sunspot.month
plot(sunspot)
library(PaleoSpec)
sp_sun <- SpecMTM(sunspot)
LPlot(sp_sun)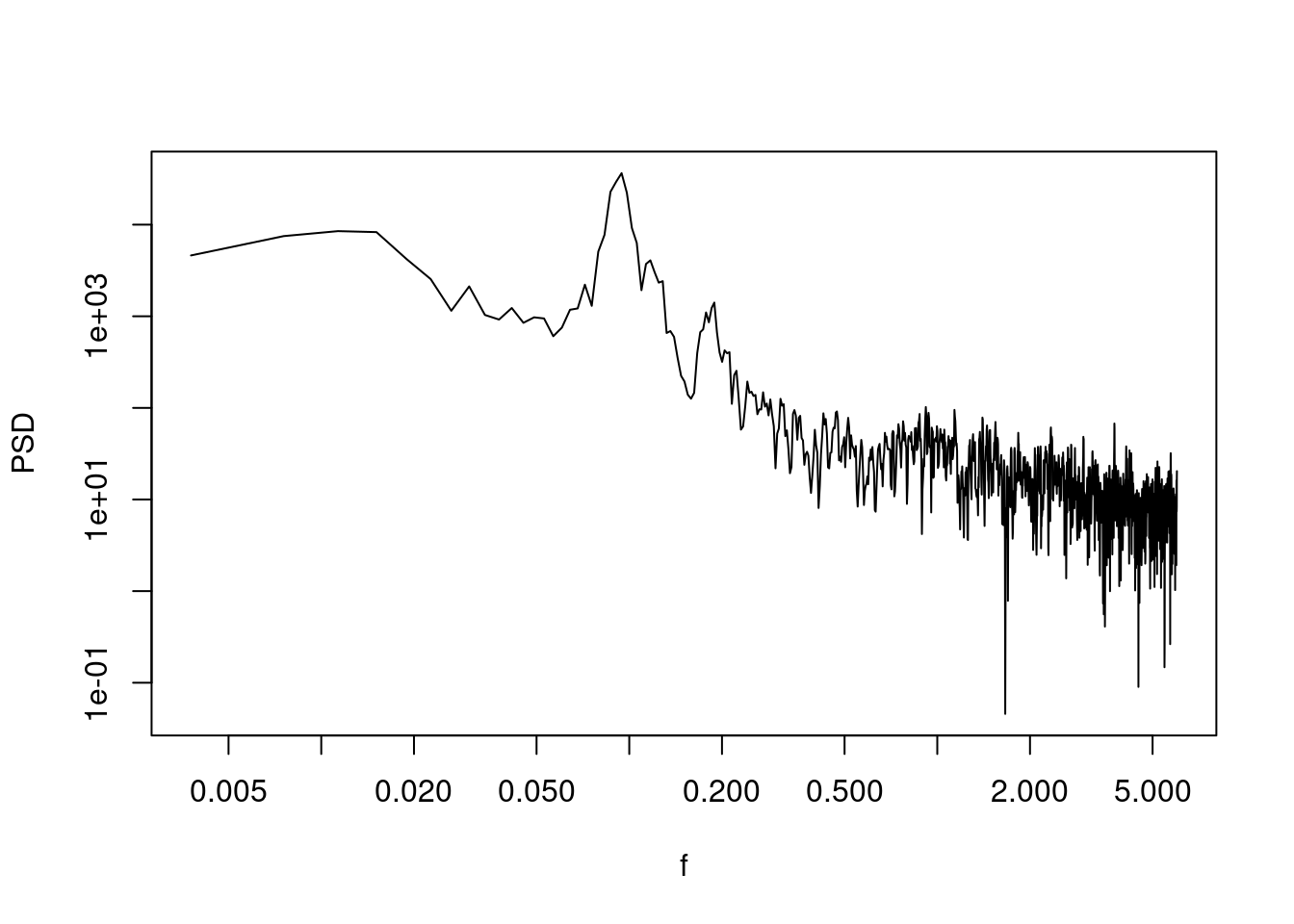
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(tidyr)
library(PaleoSpec)
library(ggplot2)
f1 <- 1/10
f2 <- 1/100
f3 <- 1/1000
tau <- 1e04
#tau / (1/c(f1, f2, f3))
dat <- tibble(
time = seq(0, tau, by = 1),
y = cos(2*pi*f1*time) + cos(2*pi*f2*time) + cos(2*pi*f3*time)
) %>%
mutate(y = y + rnorm(n(), 0, 0.1))
plot(dat$time, dat$y, type = "l")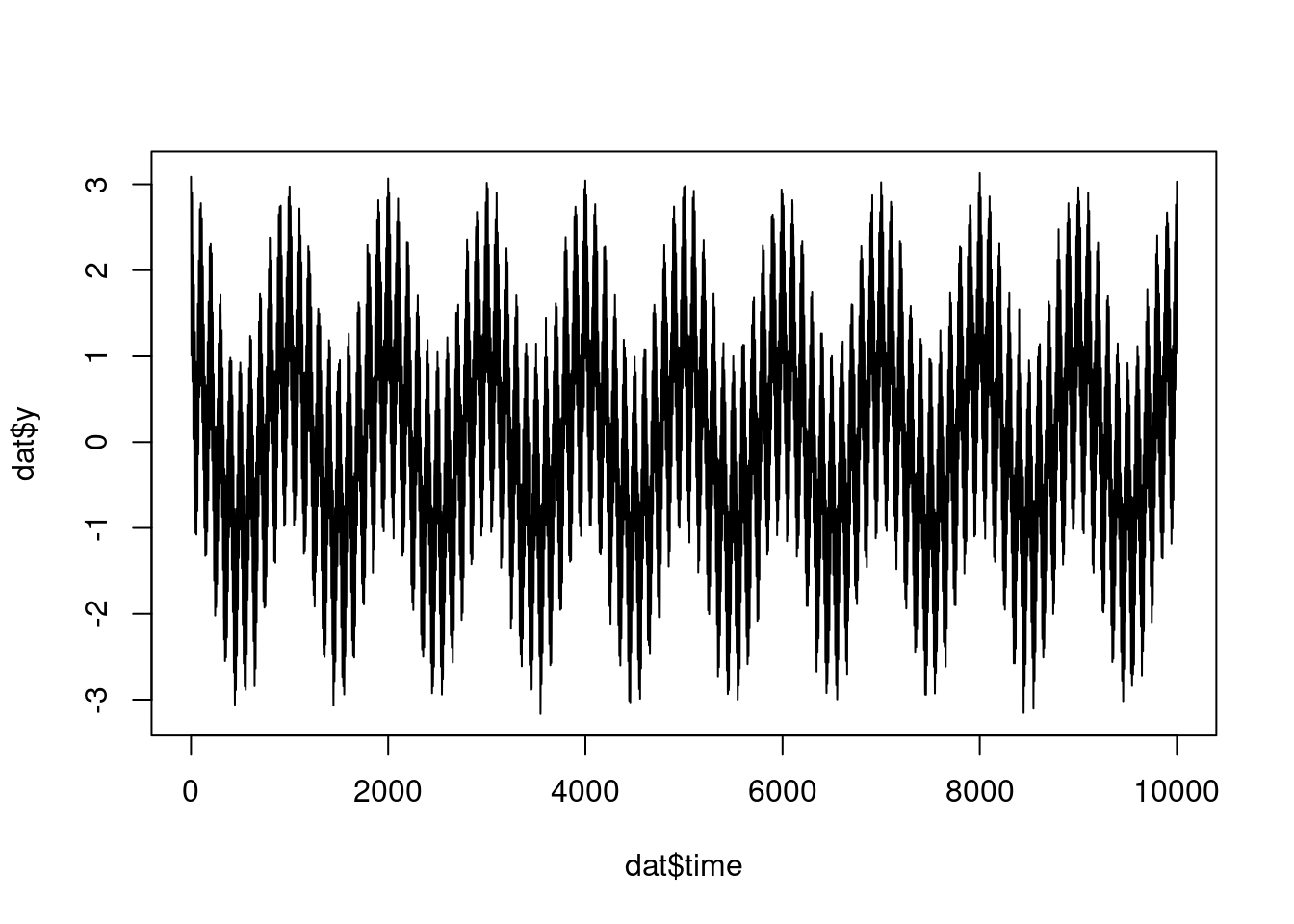
sp1 <- SpecMTM(ts(dat$y, deltat = 1), detrend = FALSE)
sp1_detr <- SpecMTM(ts(dat$y, deltat = 1), detrend = TRUE)
#sp1 <- SpecACF(ts(dat$y, deltat = 1), bin.width = 1, detrend = FALSE)
gg_spec(list(sp1, detrended = sp1_detr)) +
geom_vline(xintercept = c(f1, f2, f3), lty = 2, colour = "Red")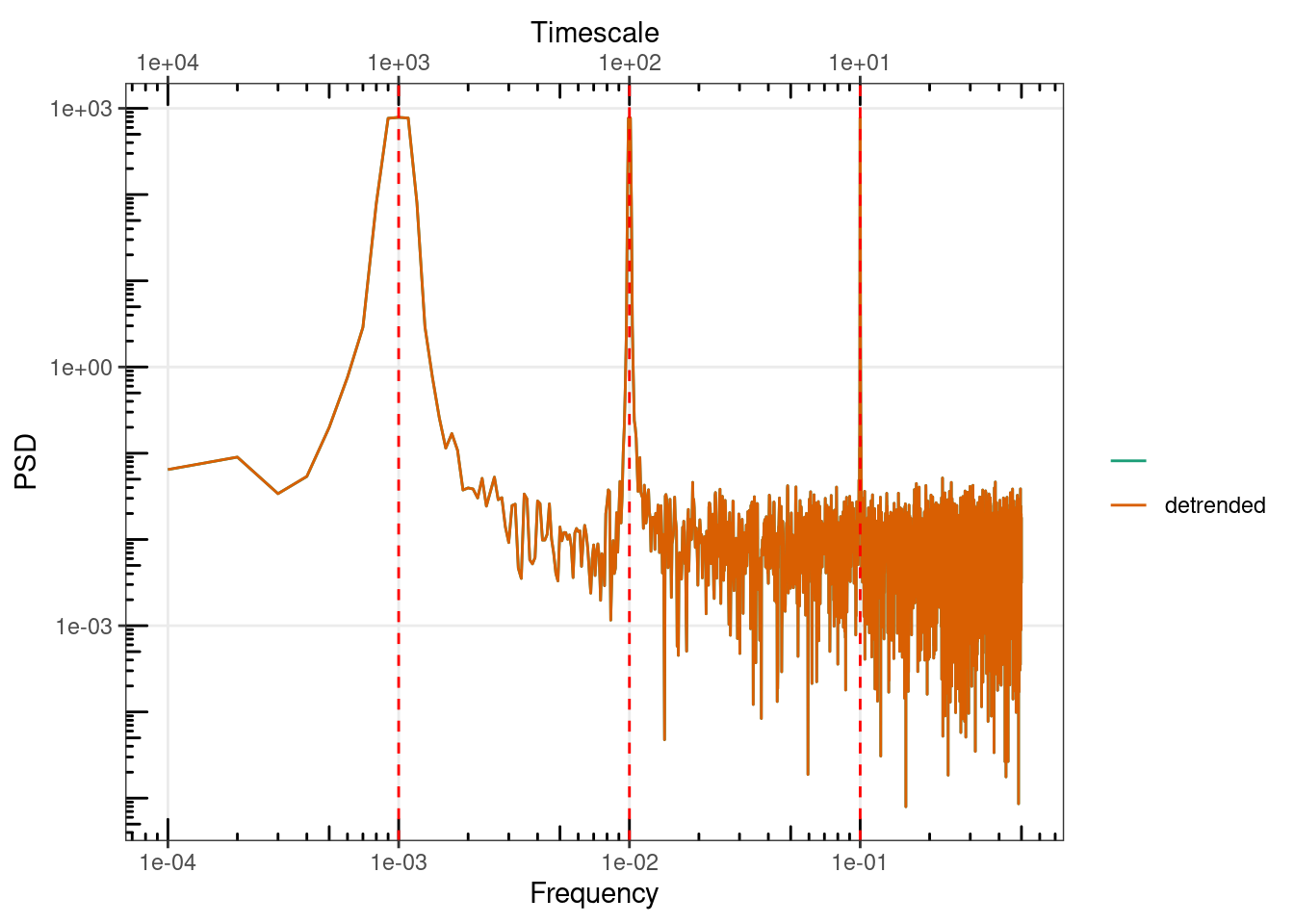
sp1_10k <- SpecACF(ts(dat$y, deltat = 1), bin.width = 1, detrend = FALSE)
sp1_10k_detr <- SpecACF(ts(dat$y, deltat = 1), bin.width = 1, detrend = TRUE)
plot(y~time, data = dat[1:500,], type = "l")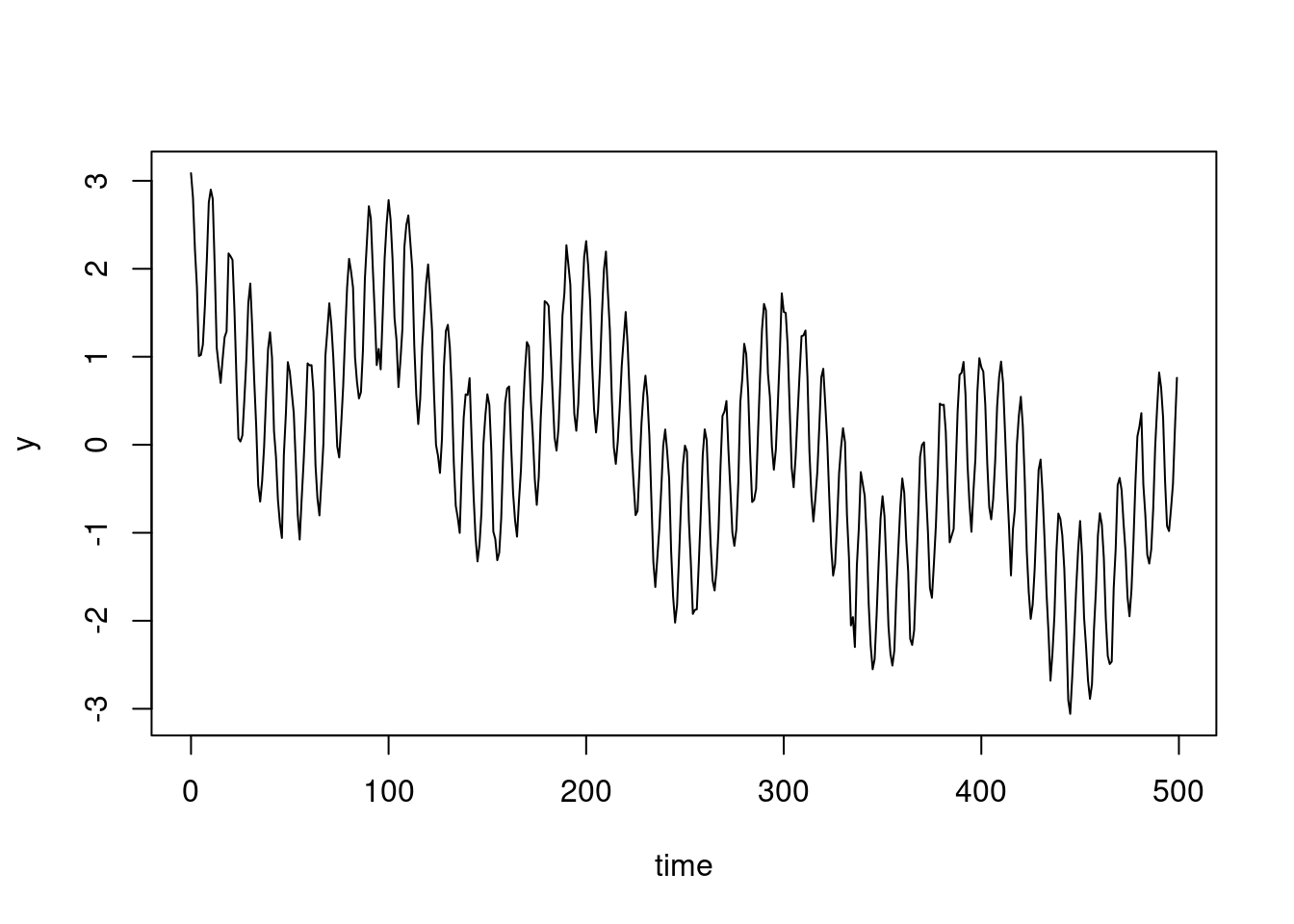
sp1_0.5k <- SpecACF(ts(dat$y[1:500], deltat = 1), bin.width = 1, detrend = FALSE)
sp1_0.5k_detr <- SpecACF(ts(dat$y[1:500], deltat = 1), bin.width = 1, detrend = TRUE)
sp1_0.5k_detr_MTM <- SpecMTM(ts(dat$y[1:500], deltat = 1), detrend = TRUE)
list(
raw_10k = (sp1_10k),
detrended_10k = (sp1_10k_detr),
raw_0.5k = (sp1_0.5k),
detrended_0.5k = (sp1_0.5k_detr)
#detrended_0.5k_MTM = (sp1_0.5k_detr_MTM)
) %>%
lapply(., Spec2DF) %>%
bind_rows(., .id = "name") %>%
separate(name, into = c("detrended", "length"), sep = "_") %>%
mutate(spec_id = detrended) %>%
as_spec_df() %>%
gg_spec(., colour = detrended) +
facet_wrap(~length)#+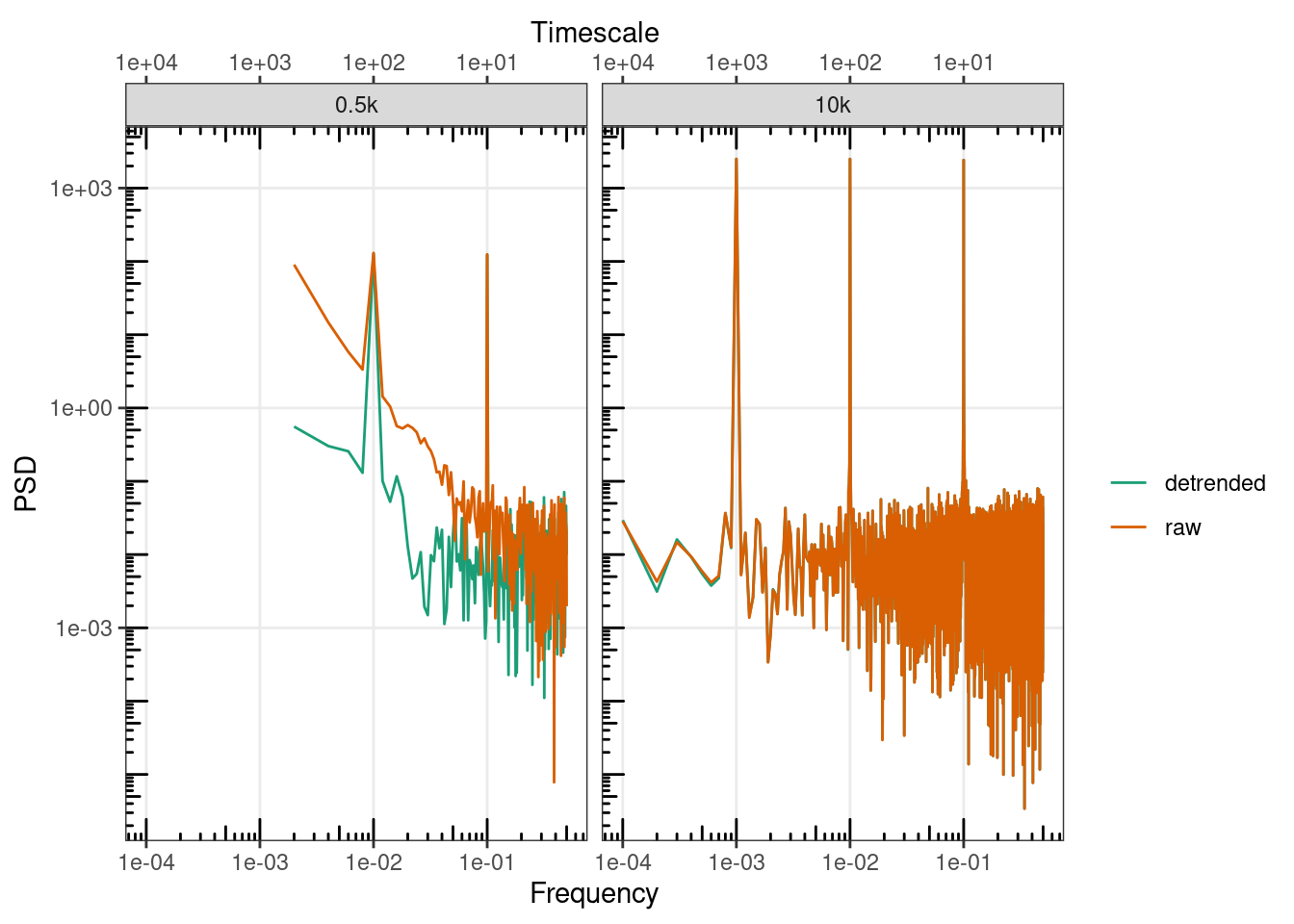
#geom_vline(xintercept = c(f1, f2, f3), lty = 2, colour = "blue")library(dplyr)
library(tidyr)
library(PaleoSpec)
library(ggplot2)
N <- 1e03
n_rep <- 1000
alpha = 0.1
beta = 0
dat <- crossing(
rep = 1:n_rep,
t = 1:N
) #%>%
# group_by(rep) %>%
# mutate(
# y = PaleoSpec::SimPLS(N = N, beta = beta, alpha = alpha)
# )
# Get a matrix of simulated timeseries
ts_m <- replicate(n_rep, PaleoSpec::SimPLS(N = N, beta = beta, alpha = alpha))
mean_spec <- SpecACF(ts_m, bin.width = 1, detrend = FALSE)
mean_spec_detrended <- SpecACF(ts_m, bin.width = 1)
dat$y <- as.numeric(ts_m)
specs_mtm_detrend <- dat %>%
group_by(rep) %>%
do({
Spec2DF(SpecMTM(ts(.$y),
detrend = TRUE
)
)
})
specs_mtm_raw <- dat %>%
group_by(rep) %>%
do({
Spec2DF(SpecMTM(ts(.$y),
detrend = FALSE
)
)
})
mean_spec_MTM_raw <- specs_mtm_raw %>%
group_by(freq) %>%
summarise_if(is.numeric, mean)
mean_spec_MTM_detrended <- specs_mtm_detrend %>%
group_by(freq) %>%
summarise_if(is.numeric, mean)
gg_spec(list("Periodogram_raw_ts" = mean_spec,
"Periodogram_detrended_ts" = mean_spec_detrended,
MTM_detrended = mean_spec_MTM_detrended,
MTM_raw_ts = mean_spec_MTM_raw)) #+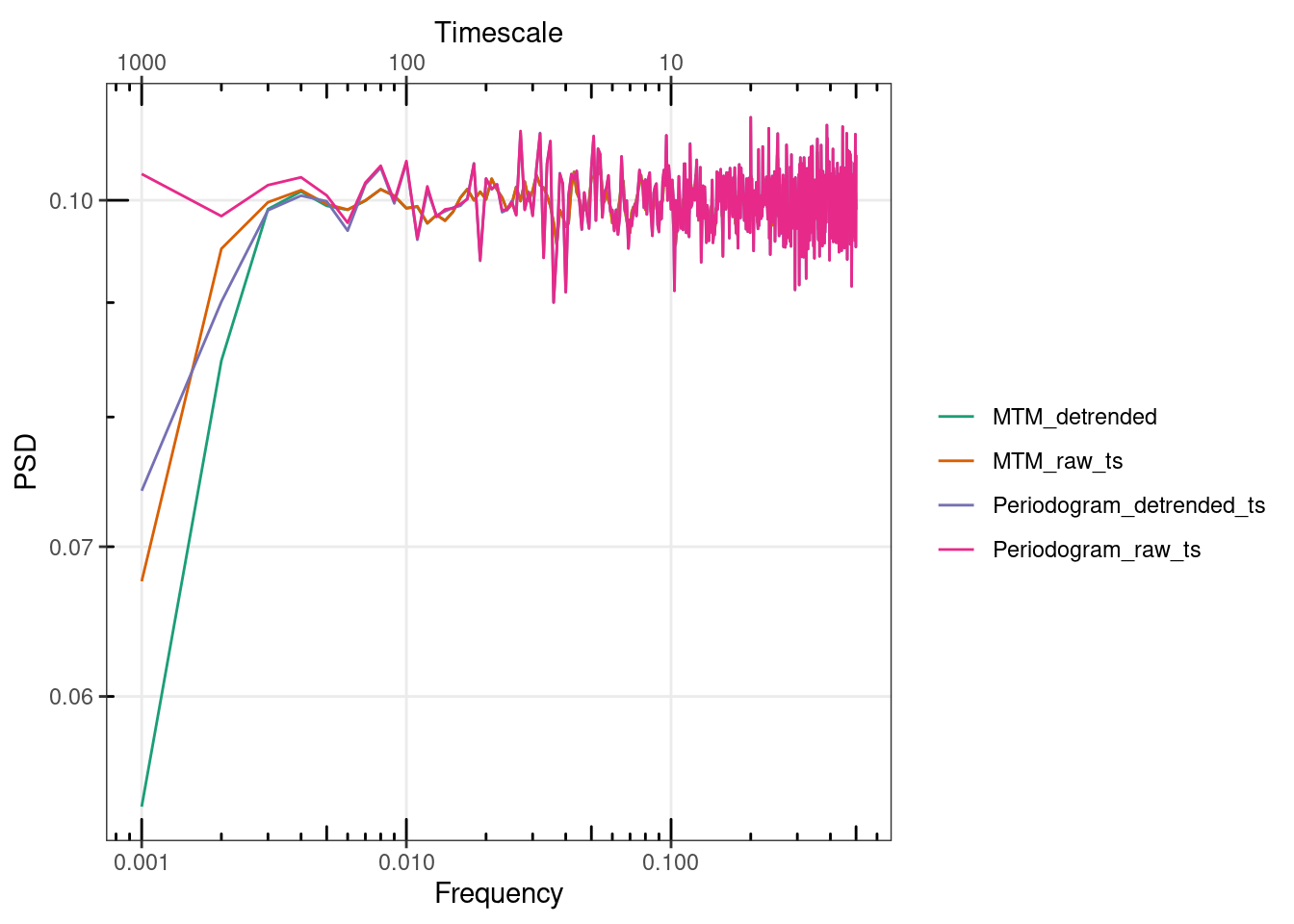
#geom_point(aes(x = freq, y = spec, colour = spec_id))